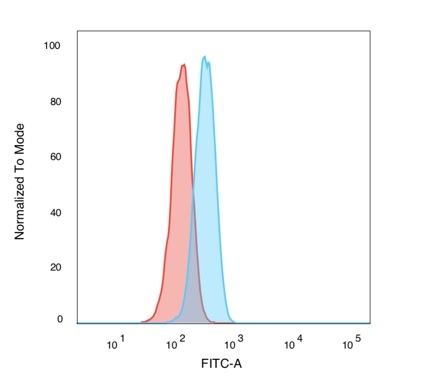
Flow Cytometric Analysis of PFA-fixed K562 cells. POLE3 Mouse Monoclonal Antibody (PCRP-POLE3-2F10) followed by goat anti-mouse IgG-CF488 (blue); isotype control (red).
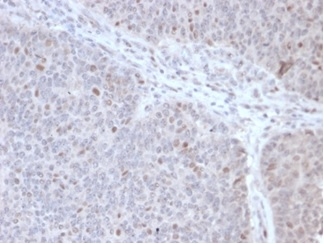
Formalin-fixed, paraffin-embedded human lymphoma stained with POLE3 Mouse Monoclonal Antibody (PCRP-POLE3-2F10) at 2ug/ml. HIER: Tris/EDTA, pH9.0, 45min. 2°C: HRP-polymer, 30min. DAB, 5min.
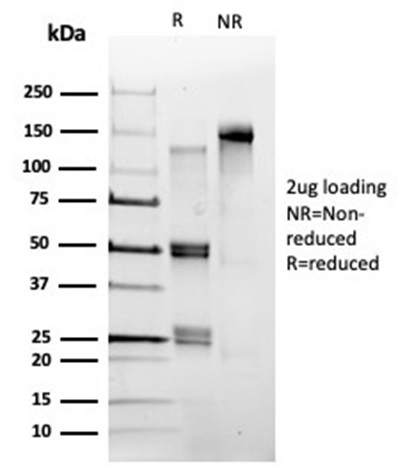
SDS-PAGE Analysis. Purified POLE3 Mouse Monoclonal Antibody (PCRP-POLE3-2F10). Confirmation of Purity and Integrity of Antibody.

Analysis of Protein Array containing more than 19,000 full-length human proteins using POLE3 Mouse Monoclonal Antibody (PCRP-POLE3-2F10). Z- and S- Score: The Z-score represents the strength of a signal that a monoclonal antibody (MAb) (in combination with a fluorescently-tagged anti-IgG secondary antibody) produces when binding to a particular protein on the HuProtTM array. Z-scores are described in units of standard deviations (SD's) above the mean value of all signals generated on that array. If targets on HuProtTM are arranged in descending order of the Z-score, the S-score is the difference (also in units of SD's) between the Z-score. S-score therefore represents the relative target specificity of a MAb to its intended target. A MAb is considered to specific to its intended target, if the MAb has an S-score of at least 2.5. For example, if a MAb binds to protein X with a Z-score of 43 and to protein Y with a Z-score of 14, then the S-score for the binding of that MAb to protein X is equal to 29.
DNA replication is initiated by the binding of initiation factors to the origin of replication. Nucleosomes inhibit access to the replication machinery at these origin sequences. Nucleosome remodeling factors increase the accessibility of nucleosomal DNA to transcriptional regulators. CHRAC15 and CHRAC17 are subunits of the nucleosomal remodeling factor CHRAC (chromatin accessibility complex), which increases the accessibility of nucleosomal DNA in an ATP-dependent manner. Unlike other known chromatin remodeling factors, CHRAC also functions during chromatin assembly by using ATP to convert irregular chromatin into a regular array of nucleosomes with even spacing. This conversion process occurs when CHRAC organizes randomly deposited histones into a regularly spaced array. In the presence of CHRAC, the nucleosomal ATPase ISWI catalyzes several ATP-dependent transitions of chromatin structure.
There are no reviews yet.