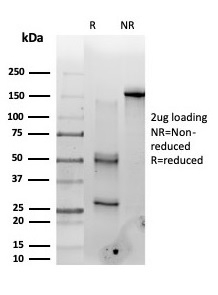
SDS-PAGE Analysis of Purified PMS1 Mouse Monoclonal Antibody (PCRP-PMS1-2E11). Confirmation of Purity and Integrity of Antibody.
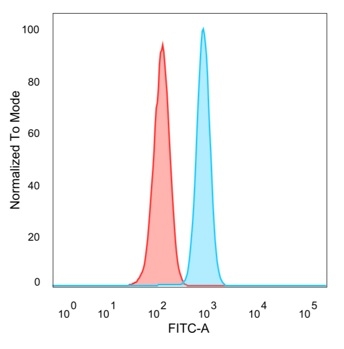
Flow cytometric analysis of PFA-fixed HeLa cells. PMS1 Mouse Monoclonal Antibody (PCRP-PMS1-2E11) followed by goat anti-mouse IgG-CF488 (blue); isotype control (red).
![Immunofluorescence Analysis of PFA-fixed HeLa cells stained using PMS1 Mouse Monoclonal Antibody (PCRP-PMS1-2E11) [CF488]. PMS1 localized to nucleoplasm and nuclear bodies. Microtubules stained with CF640R. PMS1 Antibody - Image 3](https://www.neobiotechnologies.com/wp-content/uploads/2022/03/img-5378-MSM1-P0-3.jpg)
Immunofluorescence Analysis of PFA-fixed HeLa cells stained using PMS1 Mouse Monoclonal Antibody (PCRP-PMS1-2E11) [CF488]. PMS1 localized to nucleoplasm and nuclear bodies. Microtubules stained with CF640R.
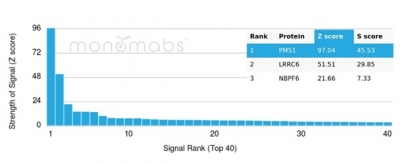
Analysis of Protein Array containing more than 19,000 full-length human proteins using PMS1-Monospecific Mouse Monoclonal Antibody (PCRP-PMS1-2E11). Z- and S- Score: The Z-score represents the strength of a signal that a monoclonal antibody (MAb) (in combination with a fluorescently-tagged anti-IgG secondary antibody) produces when binding to a particular protein on the HuProtTM array. Z-scores are described in units of standard deviations (SD's) above the mean value of all signals generated on that array. If targets on HuProtTM are arranged in descending order of the Z-score, the S-score is the difference (also in units of SD's) between the Z-score. S-score therefore represents the relative target specificity of a MAb to its intended target. A MAb is considered to specific to its intended target, if the MAb has an S-score of at least 2.5. For example, if a MAb binds to protein X with a Z-score of 43 and to protein Y with a Z-score of 14, then the S-score for the binding of that MAb to protein X is equal to 29.
The finding that mutations in DNA mismatch repair genes are associated with hereditary nonpolyposis colorectal cancer (HNPCC) has resulted in considerable interest in the understanding of the mechanism of DNA mismatch repair. Initially, inherited mutations in the MSH2 and MLH1 homologs of the bacterial DNA mismatch repair genes MutS and MutL were demonstrated at high frequency in HNPCC and were shown to be associated with microsatellite instability. The demonstration that 10 to 45% of pancreatic, gastric, breast, ovarian and small cell lung cancers also display microsatellite instability has been interpreted to suggest that DNA mismatch repair is not restricted to HNPCC tumors but is a common feature in tumor initiation or progression. Two additional homologs of the prokaryotic MutL gene, designated PMS1 and PMS2, have been identified and shown to be mutated in the germline of HNPCC patients.
There are no reviews yet.